Abstract
Introduction. ETO2-GLIS2 (aka CBFA2T3-GLIS2) is the most common (30%) alteration in pediatric de novo acute megakaryoblastic leukemia (AMKL). These patients have poor response to induction therapy, a high incidence of relapse (~90%), and dismal 5-year survival rates (<20%). Previous studies suggest that ETO2-GLIS2 induces leukemia through abnormal enhancer formation as a single oncogenic "hit". Because ETO2-GLIS2 expression induces formation of leukemia-specific neo-superenhancer (SE) elements, we hypothesize that mediator (MED) proteins are involved in linking neo-SE elements to distal gene expression, and thus could be a therapeutic target. Here, we analyzed expression of MED-family genes in ETO2-GLIS2 positive AMKL patients in combination with enhancer-histone marks, and evaluated impact of MED-kinase inhibition with selective CDK8 inhibitors (CDKi) against cell viability.
Methods. To address our hypothesis, we analyzed published RNA sequencing dataset (Smith et al. 2020) for human MED-genes expression from a pan-pediatric AML cohort (N=1476). The cohort consisted of subgroups expressing fusions of ETO2-GLIS2 (N=40), CBFB-MYH11 (N=174), DEK-NUP214 (N=49), KMT2A-ELL (N=50), KMT2A-MLLT10 (N=86), KMT2A-MLLT3 (N=114), KMT2A-MLLT4 (N=49), NUP98-NSD1 (N=107), RUNX1-RUNX1T1 (N=210) and no detectable fusions (N=526), compared to normal bone marrow (NBM) samples (N=71). Enrichment of histone marks overlapping MED-genes was analyzed from published chromatin immunoprecipitation (ChIP) sequencing in ETO2-GLIS2 positive M-07e AMKL line (Thirant et al. 2017). We tested efficacy of CDK8 inhibition with BI-1347 and CCT251545 against M-07e cells to determine their activity in the context of marked MED12L overexpression.
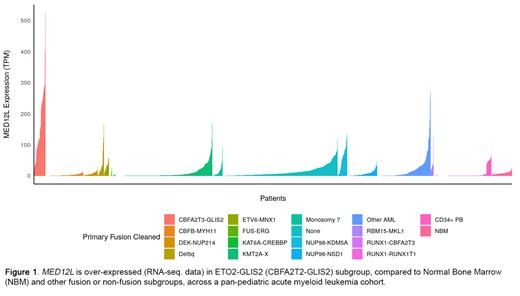
Results. We examined expression of 29 MED-genes comprising the 4 major (head, middle, tail, and kinase) MED-modules. MED gene expression was variable across AML subtypes and NBM. However, MED genes were more commonly over-expressed in the ETO2-GLIS2 group, in particular MED 17, MED1, MED10, MED27, and MED12L (paralog of MED12) were upregulated in the subgroup. Most notably, we noted exceptional upregulation of MED12L (FC 4.9, log2), compared to NBM (Figure 1). Because MED12/12L plays an intrinsic biological role in establishing oncogenic enhancer-expression loops in hematopoietic stem or leukemic cells, we investigated the overlap of enhancer bound histones such as H3K27ac and H3K4me1 to MED12L in M-07e cells, compared to umbilical cord blood-derived normal megakaryoblasts (MK) (S004BT; Blueprint epigenome database). We found enrichment of H3K4me1 at MED12L transcription start site (TSS) and upstream promoter both in MK and M-07e cells. In addition, we observed a large region of H3K27ac enrichment spanning 89 Kb (16 kb upstream and 73 kb downstream) across MED12L TSS in M-07e cells, suggesting neo-enhancer activity at this locus. Considering the dependency of MED12 on CDK8 for MED-kinase activities (Klatt et al. 2020), we treated M-07e cells with CDK8i(s), to test our hypothesis if perturbation of epigenetically enhanced MED12L expression can impact leukemic growth. However, we observed a poor correlation between M-07e cell viability and IC 50 of BI-1347 (IC 50: 0.87 µM, R 2: 0.36) or CCT251545 (IC 50: 0.4 µM, R 2: 0.48). In contrast, ETO2-GLIS2 negative MV4-11 AML cells were susceptible to both BI-1347 (IC 50: 0.44 µM, R 2: 0.88) and CCT251545 (IC 50: 0.12 µM, R 2: 0.87). Given the inefficacy of CDK8i against M-07e, and cooperativity of bromodomain extra-terminal (BET)-BRD4 and MED12/12L in forming enhancer complexes, we tested possible inhibitory impact of BRD4 inhibitor JQ1 on MED12L expression and leukemic growth in target cells. We observed effective reduction in M-07e cell viability (IC 50:0.3 µM, R 2: 0.93) with concomitant reduction not only in BRD4 protein-expression, but diminished MED12L protein expression at IC 50 and higher doses of JQ1.
Conclusion. Our findings revealed that MED12L is highly overexpressed and overlapped with strong neo-enhancer chromatin-marks in ETO2-GLIS2 positive cells, while maintaining resistance to CDK8i. Future studies will contribute to deeper insights into the preferential recruitment and role of MED12L in ETO2-GLIS2 bound enhancers and potential mechanisms of resistance to CDK8 inhibition in the disease.
No relevant conflicts of interest to declare.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal